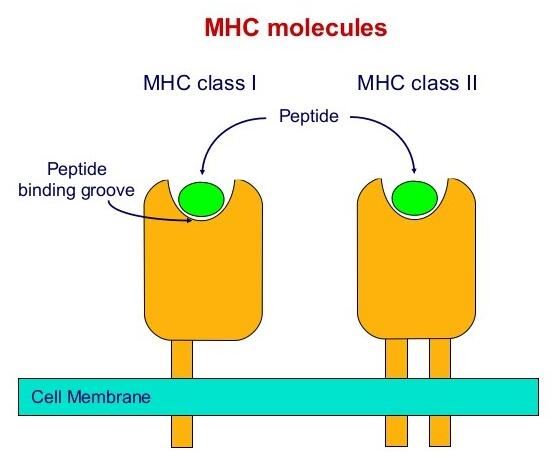
High-throughput peptide-MHC complex generation and kinetic screenings of TCRs with peptide-receptive HLA-A*02:01 molecules | Science Immunology
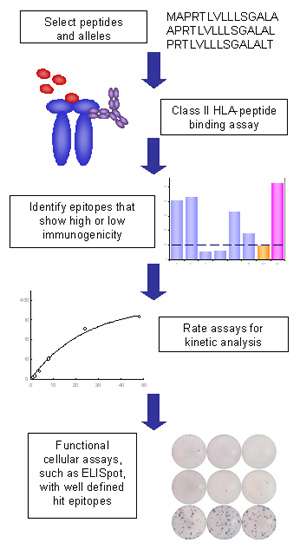
MHC Class II Binding Assays - ProImmune - Mastering Immunity _ MHC pentamers, CD1d tetramers, custom peptide synthesis, immunoassays, T cell epitope discovery, HLA tissue typing, cellular analysis, ligand binding assays, immune response

Combining Three-Dimensional Modeling with Artificial Intelligence to Increase Specificity and Precision in Peptide–MHC Binding Predictions | The Journal of Immunology
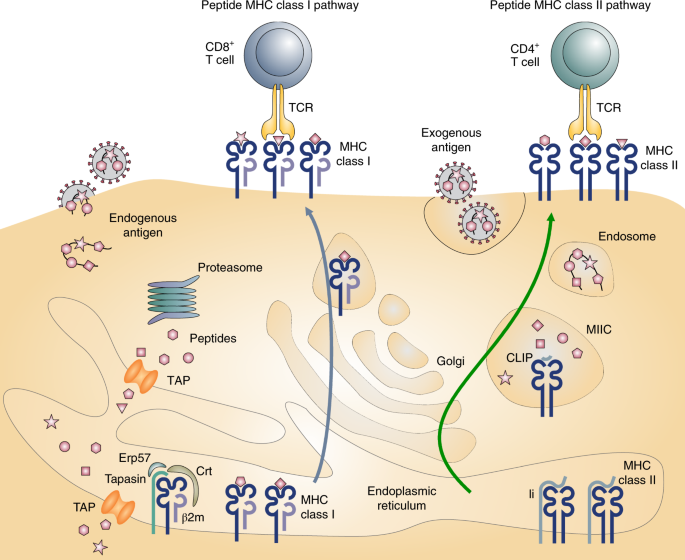
Mass spectrometry–based identification of MHC-bound peptides for immunopeptidomics | Nature Protocols

HLA-DM catalytically enhances peptide dissociation by sensing peptide–MHC class II interactions throughout the peptide-binding cleft - Journal of Biological Chemistry

Precision Neoantigen Discovery Using Large-scale Immunopeptidomes and Composite Modeling of MHC Peptide Presentation - Molecular & Cellular Proteomics
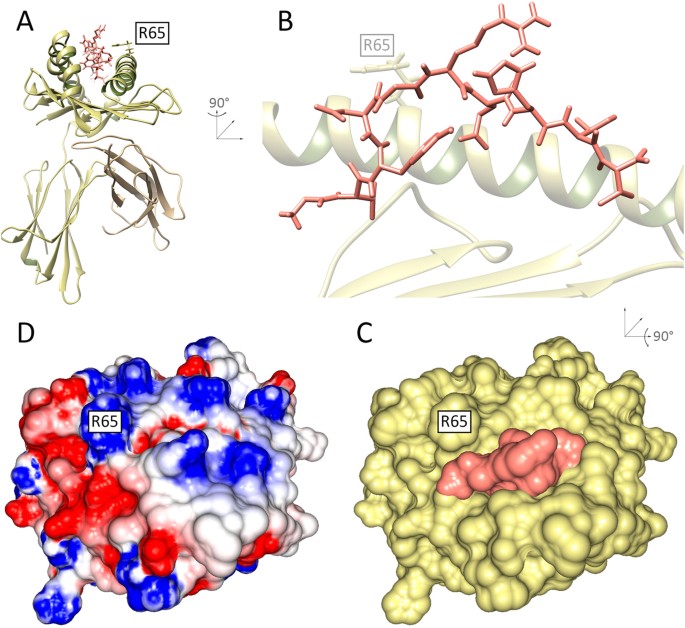
General Prediction of Peptide-MHC Binding Modes Using Incremental Docking: A Proof of Concept | Scientific Reports

MHC-peptide binding assays. MHC binding affinities are determined in... | Download Scientific Diagram

Peptide Modulation of Class I Major Histocompatibility Complex Protein Molecular Flexibility and the Implications for Immune Recognition* - Journal of Biological Chemistry

High-resolution profiling of MHC II peptide presentation capacity reveals SARS-CoV-2 CD4 T cell targets and mechanisms of immune escape | Science Advances

Repertoire-scale determination of class II MHC peptide binding via yeast display improves antigen prediction | Nature Communications

Three types of MHC binding peptides based on patient HLA allele types.... | Download Scientific Diagram
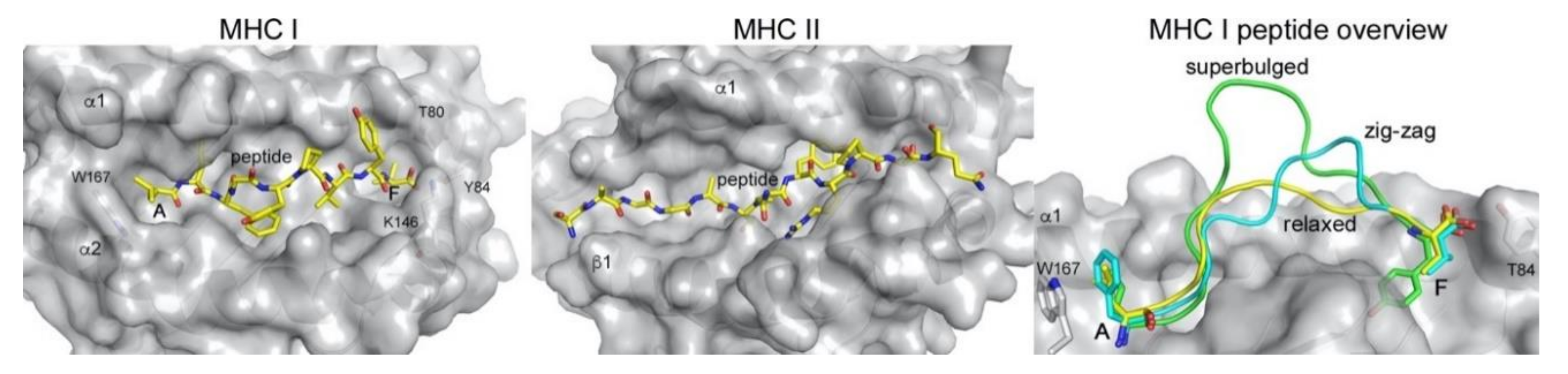
IJMS | Free Full-Text | Unconventional Peptide Presentation by Classical MHC Class I and Implications for T and NK Cell Activation | HTML

The Length Distribution of Class I–Restricted T Cell Epitopes Is Determined by Both Peptide Supply and MHC Allele–Specific Binding Preference | The Journal of Immunology

Towardmore accurate pan-specific MHC-peptide binding prediction : a review of current methods and tools | Semantic Scholar
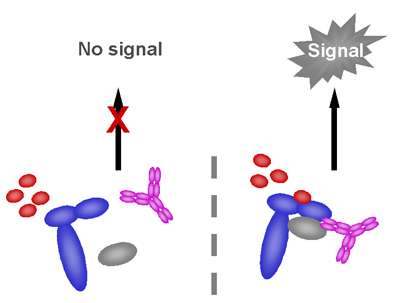
MHC Class I Binding Assays - ProImmune - Mastering Immunity _ MHC pentamers, CD1d tetramers, custom peptide synthesis, immunoassays, T cell epitope discovery, HLA tissue typing, cellular analysis, ligand binding assays, immune response
![PDF] Sensory neurons with MHC-like peptide binding properties: disease consequences. | Semantic Scholar PDF] Sensory neurons with MHC-like peptide binding properties: disease consequences. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6a6319316abdf60dbe865dc320c3bb45d82dd3c9/3-Figure1-1.png)
PDF] Sensory neurons with MHC-like peptide binding properties: disease consequences. | Semantic Scholar

T cell receptor and coreceptor CD8 alphaalpha bind peptide-MHC independently and with distinct kinetics | Davis Lab Oxford | Structure-Based Immunology