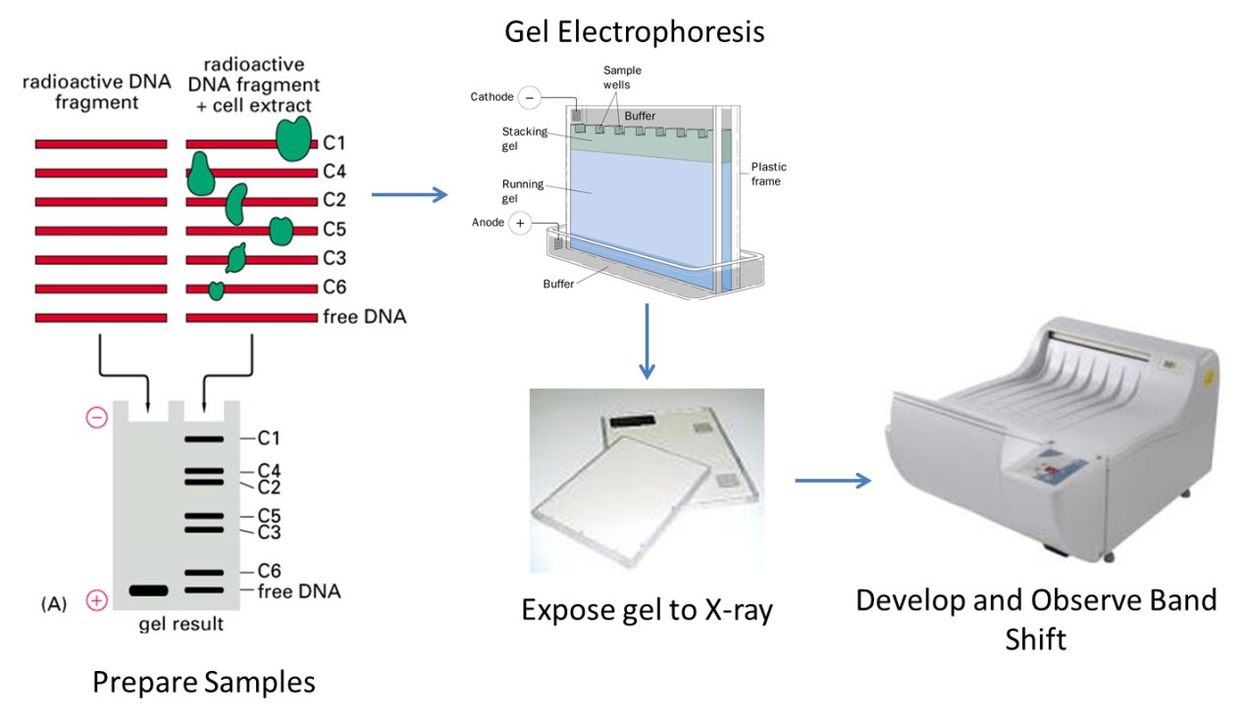
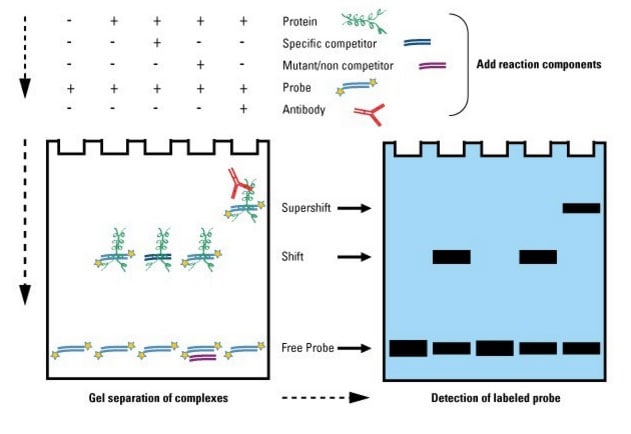
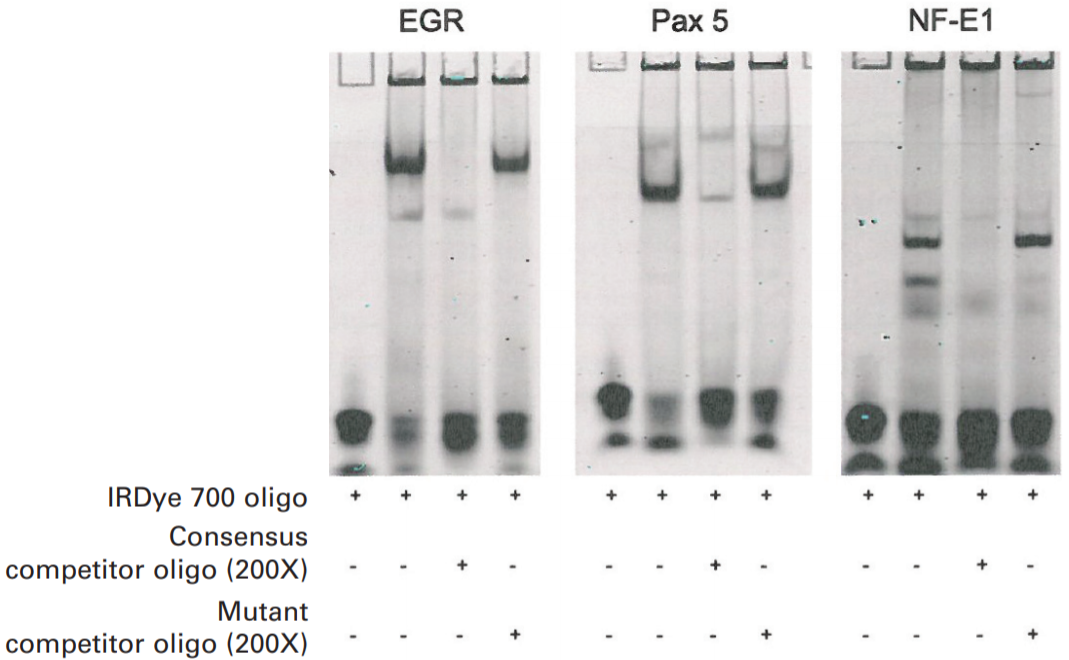
Electrophoretic mobility shift assay (EMSA) to indicate OxyR binding to the promoter region upstream of phnW.

Structural characterization of VapB46 antitoxin from Mycobacterium tuberculosis: insights into VapB46–DNA binding - Roy - 2019 - The FEBS Journal - Wiley Online Library
A Proteomics Approach for the Identification of DNA Binding Activities Observed in the Electrophoretic Mobility Shift Assay*

Electrophoretic Mobility Shift Assay (EMSA) for the Study of RNA-Protein Interactions: The IRE/IRP Example | Protocol
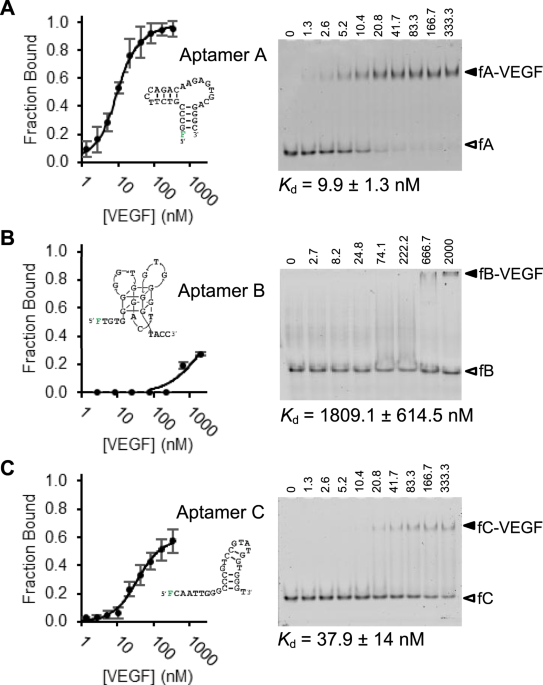
Unraveling Determinants of Affinity Enhancement in Dimeric Aptamers for a Dimeric Protein | Scientific Reports
Electrophoretic Mobility Shift Assay (EMSA) of unmodified TBA2 with... | Download Scientific Diagram
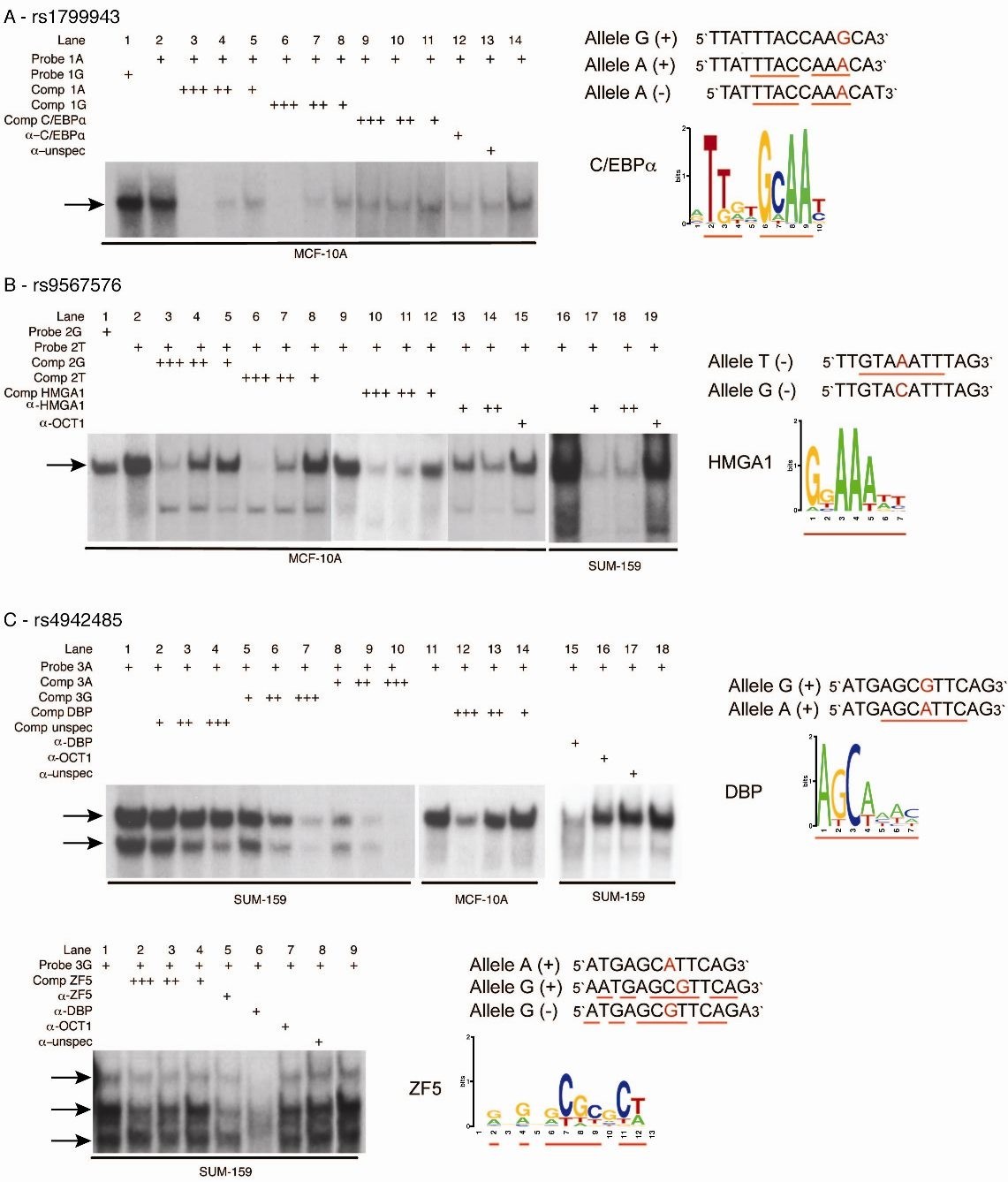
Effects of BRCA2 cis-regulation in normal breast and cancer risk amongst BRCA2 mutation carriers | Breast Cancer Research | Full Text
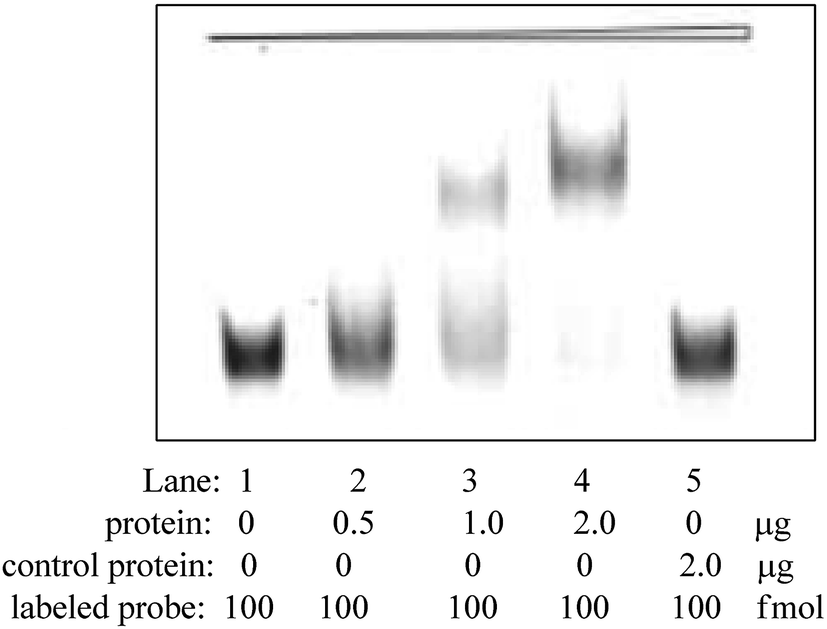
In vitro selection of DNA aptamers for the development of chemiluminescence aptasensor for neuron-specific enolase (NSE) detection - RSC Advances (RSC Publishing) DOI:10.1039/C9RA00785G
Label‐Free Electrophoretic Mobility Shift Assay (EMSA) for Measuring Dissociation Constants of Protein‐RNA Complexes

Why doesn't 100 percent band shift occur in EMSA even when binding saturation has been achieved? | ResearchGate

The Identification of Nucleic Acid-interacting Proteins Using a Simple Proteomics-based Approach That Directly Incorporates the Electrophoretic Mobility Shift Assay * - Molecular & Cellular Proteomics

Electrophoretic mobility shift assays (EMSA) showing binding of CRP to... | Download Scientific Diagram

EMSA was performed to quantitatively analyse DNA-protein interaction.... | Download Scientific Diagram

Electrophoretic Mobility Shift Assay (EMSA) for the Study of RNA-Protein Interactions: The IRE/IRP Example | Protocol

EMSA analysis of the stoichiometry of AddAB binding to dsDNA ends. (A)... | Download Scientific Diagram